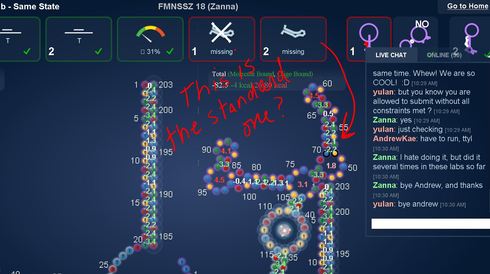
This problem has happened repeatedly to me while doing current labs.
Just to verify - is this in natural mode?
Yes. Andrew said it’s because the sequence isn’t the standard one and the program doesn’t recognize it for that reason. He said we should experiment with it anyway, though it will show constraints are not met. But it’s confusing when it’s not recognized and tells it’s not there. Anyway, I guess it’s not really a bug after all.
Scratch that comment. I JUST had it happen again with a different one. Geez, I don’t know if this is the standard one or not. This is natural mode.(it has to be in natural mode. That’s the only way to see where the hairpin is in these puzzles.)
The first one does not look like the normal MS2, the one above does. The “normal” one has 3 C’s near the tail end. You are seeing the problem regardless of which version you use. Interesting!
I read in the forums, somewhere, that there were over 20 different versions of the hairpin in a previous lab. Maybe that is still the case in this round. Not sure. Anyway, I find it confusing.
The standard one should have a blue NT at 55. Your example in the above image doesn’t so I think all is OK.
Thanks for the reply, JR. The real problem was that I didn’t realize there were multiple hairpins being used. AND that all but the standard are not recognized. All is well, it seems.
Some background: A couple of years ago Nando “slipped in” the ability for the MS2 stamper to stamp alternative sequences, at Brourd’s encouragement, with no general announcement. Nando was (and is) concerned that actually checking the puzzle constraints for each possible alternative sequence would be tricky to implement without slowing down the UI for all players.
Since then, knowledge of the capability, and its rationale for existence, has been spreading to more players. But it is still more an “experimental prototype” for the adventurous player than a supported feature. For more info, see https://getsatisfaction.com/eternagame/topics/multiple-alternatives-for-the-ms2-aptamer-sequence.
Some of the puzzles just don’t like the active MS2, by that I mean I have difficulty getting a solution I like. Since starting to use alternate MS2s I find I can find solutions for those puzzles easier.
I do it too some times, but the question is- is data gathered from comparing different types of MS2 without it being part of the experiment ok? I have absulutly no idea, not saying that it certainly bad.
I ran into the same problem early on in the pre-labs (https://getsatisfaction.com/eternagame/topics/please-review-the-proposed-puzzles-for-the-initial-ope…)
Though Omei assured me that the alternative sequences are fine to use the fact that they are not recognized as such buy the puzzle itself is frustrating. However if all it would take to solve this is to input the alternative sequences for the MS2, then we are in luck, there are only about 25 to 30 such sequences which can be stamped.
I ran into something odd while on the Lab Puzzle Review page.
Here is the Histrory, skip to the end for conclusions :
(I was on this review page: http://www.eternagame.org/game/browse/8047255/). I was zooming out to see the entire sequence at once and noticed that the associated non-sequence text failed to render for the lower puzzles while zoomed out from 80% through 50% [*Note that (for my computer at least) one has to zoom out to 67% in order to see entire RNA sequence]. As seen in this screenshot:
https://prntscr.com/gjsjza
Oddly enough the text appeared again when zoomed out to 33%. I tried a different puzzle (http://www.eternagame.org/game/browse/8038691/) and ran into the same issue with text vanishing while zoomed out, BUT at DIFFERENT percentages of zoom.
I reloaded the first page and noticed that everything appeared fine when zoomed out until I noticed that the problem was still there, but now the lower portion of the RNA sequences were gone. I continued to fuss around for a while, but then finally stumbled upon the reason for the anomaly.
Conclusion :
The RNA sequences scale at a different rate relative to the rest of the text in the review page. Thus, it appears that text fails to render for either sequence or design details, but in reality the information is there, but is offset by zooming in or out.
Why this is an Issue :
If I zoom out in order to look at my sequences, then it is likely that the sequences immediately to the right of my design details are not actually my sequences, but rather those of the submissions a few ahead or a few behind me, with the disparity increasing with the among (percent) of zoom. With zooming in/out the only way to know which sequence goes where is to find the unique features in one’s RNA sequence when the screen is at 100% and then to keep track of that sequence as one zooms out to see where it has moved to.
P.S. While I know there is a general search feature for each column, for the RNA sequences at least, it would be nice to be able to see (via highlighting as per standard “Find” search on Browsers) where in the long string of letters the exact sequence (which was searched for) is located, e.g., the aptameric sequence.