Hi all, thanks again for the perceptive questions.
The list of numbers are the data themselves, except the first number. The first number in the list is a sequence offset – we don’t always get data for the first few nucleotides for technical reasons, and that number defines the first nucleotide for which an experimental value was obtained. (or maybe the first nucleotide - 1 –> Jee can you confirm?)
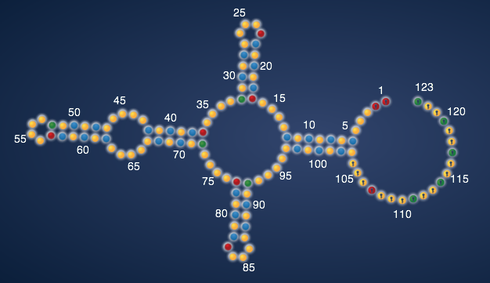
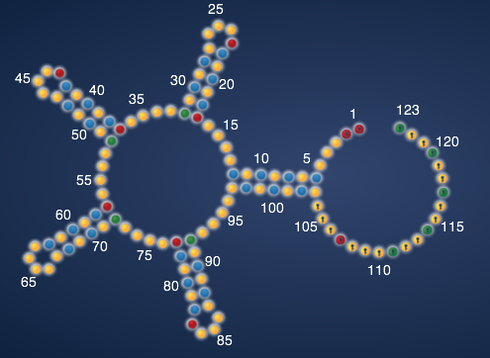
The data are the chemical accessibilities of each RNA nucleotide to the ‘SHAPE’ reagent, developed by the Weeks lab at UNC. For the aficionados, the reaction is an acylation of the 2’ hydroxyl of the RNA nucleotide by a reactive anhydride. Strongly modifed nucleotides, at least empirically, correspond well to ‘flexible’ regions of the RNA, i.e. the ‘unpaired’ parts of your EteRNA designs, if they fold correctly.
The values (min,max, and threshold) define the coloring that you see in the EteRNA viewer (threshold is white, anything at min or below is blue, and anything at max or above is yellow). These also are parameters in determining the EteRNA score:
(A) for regions that should be base paired: you get a point if the score is less than the threshold.
(B) for regions that should be unpaired: you get a point if the score is above a fairly low cutoff given by [(1/4) threshold + (3/4) min].
[We should probably make a graphic of this.] For the EteRNA score, the points are then divided to the maximum number of points (the number of nucleotides for which there is data), and then multiplied by 100.
The score was designed to be fairly accommodating but still correlate well with the designs whose SHAPE data ‘looked’ good. We are also further checking this score on several natural RNAs with known structure (stay posted on this).
The last point is the setting of min, max, and threshold – we actually optimize these 3 parameters for each design’s data via a linear program so as to give the design the maximum possible score. Again, we’re trying to give everyone the benefit of the doubt.
For the curious, we’ll soon have the data in text file format, as well as the actual raw electropherograms (which are quite beautiful, in my opinion) on a publically available website that our lab just started:
http://rmdb.stanford.edu
Stay tuned – the pipeline from EteRNA should start in a couple weeks.