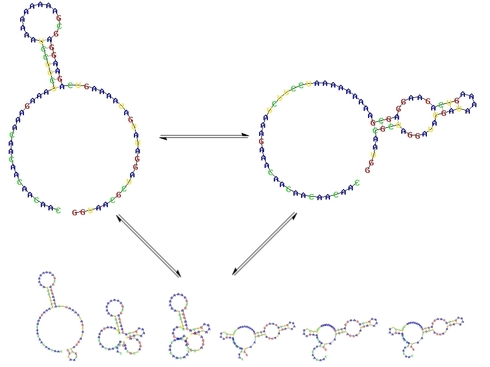
An RNA in solution shifts between many structures, with lower-energy structures more abundant at equilibrium. Certain programs can provide insight into the structure of these “suboptimal” folds and predict their abundance. This might be of particular interest in the new switch labs, since the FMN-binding conformation is one of these suboptimal structures.
RNAshapes is a versatile program that does suboptimal folding (among quite a few other things). You can submit your designs and view the output on the web.
Submitting a sequence
Sequences must be submitted in FASTA format, which looks like this:
\>sequencename
sequence
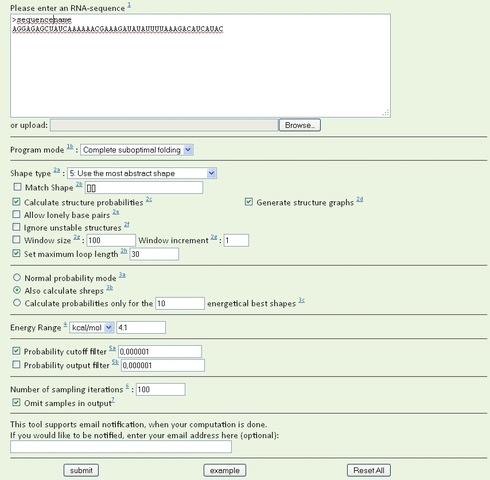
RNAshapes has many settings, most of which can be left at their defaults. You should select the following options, however:
-
Program mode - Select “complete suboptimal folding”.
-
Calculate structure probabilities - Enable.
-
Generate structure graphs - Enable.
-
Allow lonely base pairs - Enable if isolated base pairs are present in the target structure. Otherwise, optional.
-
Energy range - Select kcal/mol. If you are using the program in a switch lab, you must use a value greater than the “energy bonus” for the design (in the first lab, this is 2.8 kcal/mol). I use a value of 4.1 kcal, which should generate all structures with at least 1/1000 the abundance of the MFE structure and 1/10 the abundance of the least stable switch design you could submit. Larger energy ranges will generate more structures, but run times will be longer and the extra structures generated will have very low probabilities because of their higher energy.
Output
For ~120 nt EteRNA labs using the settings in the example above, results are ready in a minute or so. Select “results with RNA plots” from the results page to open the list of the suboptimal structures. Here are the first few lines of a sample sequence, with explanations of each column:

Interpretation
Aptamer and switch labs are new, and RNAshapes is untested as far as EteRNA goes. Moreover, suboptimal folding programs do not tell you how long a group of freshly-synthesized RNAs might take to reach equilibrium, or how quickly any two given shapes may interconvert. So why use this program? It does let us look at our designs in a different way, it may end up proving itself useful, and it points at some important questions.
If you are designing an RNA switch, do you want the two conformations to be much lower in energy than the rest of the ensemble?
Is there an ideal energy difference between the switch conformations?
How many low-energy suboptimal structures still contain the FMN-binding 6-5 loop and contribute to the FMN-binding behavior of the RNA?