Hyphema came up with a great idea of having players highlight sequences they think will be promising for mutation so other players with less time for picking out designs will have good starter places to aim their mutation spree at. So here it comes. I start this new post so we can put up what we think will be interesting mutating.
My favorite RO candidate: http://www.eternagame.org/game/solution/6892307/6975137/copyandview/
It isn’t the highest scoring, but I think it holds a higher potential for hitting a better fold change than the highscorer. Here is why.
KDOFF/KDON = Foldchange
KDOFF: 59
KDON: 4, 4, 13 (Worst) - 13 is for the B input. It takes more for it to bind. But if this number get lower, foldchange will raise.
59/13 = 4.64
Foldchange potential
59/4 = 14,75
Whereas the topscorer has higher KDON in general and as such doesn’t hold such a promise of lowering raising foldchange. Even if you get B binding better - which general is what is needed to get done in the RO lab, the other KDON’s are higher (7, 10 and 11 (B)) So if you get B lowered, the next worst KDON value (10) will get picked as KDON instead. However it do has a higher KDOFF (100, so its foldchange potential if next best KDON value 10 is picked is, 100/10= 10.
So this design has a higher fold change potential, if you can get the B input bind better to the design. This can be done several ways, either by strengthen complementarity to B input or and/or weakening the sequence binding up with the B complement.
Also I would like to see the B complement slided either closer or further from the rest of the puzzle. Similar slide elements in relation to each other.
Thanks for getting this started, Eli.
This definitely looks like a good example of a design where mods could make significant improvements. Here’s another idea for adjusting the KD values that seems like it could be fruitful.
I have marked four bases, numbers 34 and 44-46, that surround the reporter complement. Experiments from the first round indicate that eliminating these four bases (for example, by sliding the 5’ end of the design one base to the right, and the 3’ end three bases to the left) will decrease KD for all of the first three states. Furthermore, if the hairpin in state 4 holds, this mod should increase the KD for that state, because the smaller the hairpin, the more 3D conflict there is between the formation of a helix with the reporter and maintaining the hairpin. In fact, I’m pretty sure that making these changes would also require the hairpin stem to be strengthened.
For anyone trying this, you need to keep in mind that when it comes to stacks getting close, NUPACK predicts that free energy increases, instead of decreases. Furthermore, NUPACK can’t even fathom that in state 4, it is possible for the reporter to bind at the same time the hairpin stem holds. So we’re really on our own for predicting the effect of these changes. Expect NUPACK to predict that states 1-3 will go unstable as the stacks are made flush. And I’m not sure how to even try to compensate for NUPACK’s shortcomings in state 4.
Bottom line – when it comes to optimizing a promising design, don’t expect NUPACK to be very helpful in distinguishing between good and bad mods…
Thanks Eli for getting this started. Its getting tough for me to stick with this. maybe when my kids are older I’ll be better. However, I realize I need a new computer with these new labs. It is terribly slow for me to look at results on the archive. Then to do a puzzle it is atrociously slow. The freeze function is brilliant but I still have yet to do a lab because of the length of time it takes. So if I can click on your links here and do a puzzle then it should be better. Hopefully. If this is what we need to do then so be it. I do hope the Devs can come up with a way to integrate this “hotlink” to a lab puzzle that is promising for further “refining” into the lab pages. I think it would challenge those players that see it to give a try. If they are like me and try to look at the archives they will get frustrated with waiting to see the results download.
Hi Hyphema!
A few shortcuts for making easier mutation and still make interesting submissions.
Pick the topscorers in whatever lab. Click freeze - you can actually submit when in freeze mod and then just go on mutating without unfreezing - thus saving a ton of calculations for the hard labs.
Here are a list of mutations that I would find worth seeing on already well scoring designs.
1: Mutations of the first two bases around the reporter complement
2: Mutations of U to C and reverse for bases in the input complements
3: Mutations of G to A and reverse for bases in the input complements
4: Mutations around the inputs
5: Slides of the elements in relation to each other. Especially the reporter. (Relevant to Omei’s coaxial stacking experiment)
Especially the first one I count very interesting. It will help us figure how the flourescent tag effect our experiments + it can have great effect on the binding of the reporter and thus the score.
Here are a bunch of A*B/C^2-INC designs that I’m working with modifying:
7368799,GAGUUUUACUCUACUAGUUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAAUAGGUGUUGUGCUCGGUUCAUUUAGCCCAAAAU,A*B/C*C-INC#21,Malcolm
7368658,UUUUUUUACUAGUUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAUAAGAUGUUGUGCUCGGUUCAUUUAGCCCAAAAUUAAAU,A*B/C*C-INC#14,Malcolm
7368624,UUUUUUUACUAGUUAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAUAAGAUGUUGUGCUCGGUUCAUUUAGCCCAAAAUUAAAU,A*B/C*C-INC#13,Malcolm
7368053,GAGCAAAGCUCUACUAGAUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAAUAGAUGUUGUGCUCGGUUCAUUUAGCCCAAAAU,A*B/C*C-INC#07,Malcolm
7368028,GAGUUUUACUCUACUAGAUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAAUAGAUGUUGUGCUCGGUUCAUUUAGCCCAAAAU,A*B/C*C-INC#06,Malcolm
7367976,GAGUUUUACUCUACUAGAUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAAUAGGUGUUGUGCUCGGUUCAUUUAGCCCAAAAU,A*B/C*C-INC#05,Malcolm
7356541,GCGUUUCGCUUACUAGUUAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAAAAGAUGUUGUGCUCGGUUCAAUUAGCCCAAAAUU,jl-abc2inc-3 dl2007 mod 7072846 A55+A72,jandersonlee
7356535,GCGUUUCGCUUACUAGUUAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAAAAGAUGUUGUGCUCGGUUCAUUUAGCCCAGAAUU,jl-abc2inc-3 dl2007 mod 7072846 A55+G81,jandersonlee
7356530,GCGUUUCGCUUACUAGUUAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAAAAGAUGUUGUGCUCGGUUCAUUUAGCCCAAGAUU,jl-abc2inc-3 dl2007 mod 7072846 A55+G82,jandersonlee
7363223,UUUUUUUACUAGUGAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAAAAGGUGUUGUGCUCGGUUCUUUUAGCCCAAAAUUAAAU,ABC-INC_Modtop10_Sub002,JR
These are all fairly similar, so you could pick one or two and focus on them.
For AB/C^2-DEC here are some of the top scoring designs:
The colors in the AvsB column indicate when A or B is more reactive. Five of these (pink and red) react much more strongly to A (KD1) than to B (KD2). If we can mute the response to oglio A for these, we might improve the designs. For all of them, a stronger C response may help.
thanks Eli. I have some mods of the sequence you provided above http://www.eternagame.org/game/solution/6892307/6975137/copyandview/ but when I tried to submit I got a “you can only submit this paper lab after it is activated” message. is this not an active lab? thanks
@ Hyphema: Yes, this was from the Open TB This is it Lab started on May 1, 2016. It is closed now. The current lab is accessed from the home page, Round 3.
jl’s post below has references to current AB/C-INC lab puzzles.
jandersonlee, in the sequences for the A*B/C^2INC you gave is there something secific i need to improve on or do i just post something modified from the original sequence. thanks.
Take 7368799,GAGUUUUACUCUACUAGUUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAAUAGGUGUUGUGCUCGGUUCAUUUAGCCCAAAAU,A*B/C*C-INC#21,Malcolm for instance. My NuPACK modeling suggests that it is slightly stronger in response to A than B, but not by much (pretty well balanced, actually), and not saturated in response to either. The overall response also seems weak, so I’d probably try and make it bind more strongly to A, B, AND C. Most likely switching GCs and AUs to GUs will *not* improve this design, but I don’t know that for sure.
7368658,UUUUUUUACUAGUUAGGUGGCCCAGAAGGCGCCAGAUCAGAACUUAUAAGAUGUUGUGCUCGGUUCAUUUAGCCCAAAAUUAAAU,A*B/C*C-INC#14,Malcolm is predicted to have a stronger overall response to all oglios, but tighter or looser versions possibly might improve things.
7356541,GCGUUUCGCUUACUAGUUAGGUGGCCCAAAAGGCGCCAGAUCAGAACUUAAAAGAUGUUGUGCUCGGUUCAAUUAGCCCAAAAUU,jl-abc2inc-3 dl2007 mod 7072846 A55+A72,jandersonlee also looks fairly well balanced in the NuPACK models, but models can be off and tweaks may improve it.
However, my modelling didn’t work out spot-on in the first round, so I cannot be sure what is the best thing to tweak for any of them. So, I guess “just post something modified from the original sequence”.
Some things I will occasionally try are to “clean up the dot plot” a bit, especially around “static” stems or unbonded dangles that should appear in both states. Whether or not this is wise, I don’t know.
I find that the switch graph results imply my AB/C_INC score 60 submission looks more like the top AB/C_DEC winners from last round, so will use similar AB/C_INC submissions to look for viable AB/C_DEC winners. See graphs below.
Omei pointed out an ABC2INC design that had all its concentrations switching in the right direction while still being weak in switch.
I have made modifications of it, some of them with more reporters. I would love to see more mutants of these designs.
Single reporter variants:
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7395150&filter1_arg2=7395150&fil…
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7393490&filter1_arg2=7393490&fil…
Double middle reporter variants:
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7395637&filter1_arg2=7395637&fil…
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7395619&filter1_arg2=7395619&fil…
Tripple reporter variants:
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7395078&filter1_arg2=7395078&fil…
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7394877&filter1_arg2=7394877&fil…
http://www.eternagame.org/game/browse/7254825/?filter1_arg1=7394813&filter1_arg2=7394813&fil…
You can also help by mutating the original puzzle. Go double mutants or more. I have done a full single base mutation round on it.
http://www.eternagame.org/game/solution/6892317/7001278/copyandview/
(For background and more about the why behind, see this post)
Since I think that deleting bases to minimize or eliminate the gap between stacks is as important for improving designs as base mutations, I’ve created an EternaScript to make it easier to generate these sequences.
For motivation, consider this design of Eli’s
NUPACK gives its seal of approval to this design, but NUPACK will mispredict the MFE as stacks come together. So the idea is to start with a design that looks reasonable and NUPACK evaluates as being good, and then make incremental changes, knowing (and accepting) that these changes will cause NUPACK to mis-evaluate the MFE.
I have marked all the bases that I think are very worthy of consideration for deletion.
- Base 65 (marked in red) is one that I can practically guarantee will improve the design’s fold change if NUPACK’s estimate for the unmodified version is reasonably accurate. Making the first two hairpins in the OFF states adjacent will significantly raise KDOFF. In the ON state (state 4), reducing the size of the exterior loop should have little effect.
- Eliminating base 50 (orange) will also increase KDOFF. But it will also create a 1-bulge in the B helix of the ON state. That will have the effect of raising KDON as well, so it’s much less a sure thing that there will be a net improvement.
- Eliminating bases 41 and 42 (green) looks like another excellent bet. In the OFF state, this will reinforce both the B and R helices, which should lower KDON significantly. Plus, in the OFF state it should make it much more difficult for the reporter to bind to the hairpin loop, raising KDOFF. (Having an oligo bind to a hairpin loop while maintaining the hairpin stem is another situation that NUPACK can’t handle, but it has been widely demonstrated in the lab.)
- Eliminating base 30 (magenta) will reinforce both the A and R binding in the ON state, again lowering KD_ON. But it will also weaken the hairpin in the OFF state, lowering KD_OFF. Here, the effect on the fold change is much less clear.
Keep in mind that NUPACK is almost certain to tell you that any of these deletions will break the folding constraints. But you can still have NUPACK fold it and see whether the only states that change folding are the ones where you were purposely trying to lower the energy, knowing that NUPACK was going to think the deletion would raise it. If that happens, it is actually a very good sign. On the other hand, if both ON and OFF state foldings are changed by deleting a base, that’s an indication that some other factors are in play as well.
Great that you wrote a script to aid in deletions type mutations!
I have found a simple search space technique of aligning two sequences that fully pass all four states and then noting the differences between them and mutating them allows good discovery of sequences that also pass all four states and highlights dotplot types that become evident as potential clues and sources for development, ie.
whbob_AB/C-DEC_R2_V1.0(whbob)
AAUUUCGAACUAGGAUGGCACAAUGUAAUCAGAACUUAGG…
| | | | | | | | | | etc.
AAUUUCGAACUAGGUUGGCGCAGAUCCAGAUGAAUUUAGG…
7396629(JRmod-sub24)_29G68C
choose one form and substitute nt’s from the other 1,2,3 or more at a time to explore the space between the two forms and get new ideas. Get a promising form and use it as an offshoot for more explorations.
A script would beat my manual method (hint).
This is super awesome!
I tested your script on a concrete design where I picked two bases for deletion.
Base 42 and base 66
Script input:
Give the script a while to run. Especially if you have a lot of base deletions.
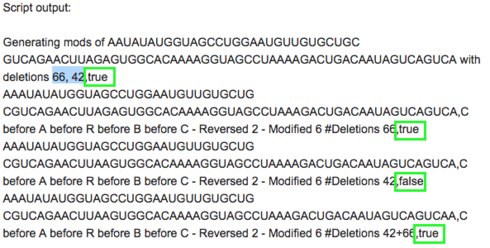
Script output:
Generating mods of AAUAUAUGGUAGCCUGGAAUGUUGUGCUGCGUCAGAACUUAGAGUGGCACAAAAGGUAGCCUAAAAGACUGACAAUAGUCAGUCA with deletions 66, 42,true
AAAUAUAUGGUAGCCUGGAAUGUUGUGCUGCGUCAGAACUUAGAGUGGCACAAAAGGUAGCCUAAAGACUGACAAUAGUCAGUCA,C before A before R before B before C - Reversed 2 - Modified 6 #Deletions 66,true
AAAUAUAUGGUAGCCUGGAAUGUUGUGCUGCGUCAGAACUUAAGUGGCACAAAAGGUAGCCUAAAAGACUGACAAUAGUCAGUCA,C before A before R before B before C - Reversed 2 - Modified 6 #Deletions 42,false
AAAUAUAUGGUAGCCUGGAAUGUUGUGCUGCGUCAGAACUUAAGUGGCACAAAAGGUAGCCUAAAGACUGACAAUAGUCAGUCAA,C before A before R before B before C - Reversed 2 - Modified 6 #Deletions 42+66,true
Jandersonlee made a script based on an earlier Omei script, that is practically perfect together with this one, as it allows to upload the script output in the game interface to view the designs directly and just click back and forth through them. (With Prev and Next).
This is saving a ton of cutting and pasting.
The script can even tell if the design is stable or not. Which is a pretty big improvement. To delete a design you don’t wish to keep and submit, just click Prune. Afterwards you can copy out the sequences you decided to keep for submission, from the bulkload window.
To use jandersonlee’s Bulk load script you need to make a copy of it and save it as a booster. You also need to change line 6, so it reflects the number in the link of your script copy. (Edit your script and update.)
Thanks, Eli.
I am a little puzzled by your output example:
I can’t figure out where the last true/false value came from. Is this perhaps actually the output after jandersonlee’s booster processes the output from my script?
Yes. When you process the lines through the bulk loader, it tries them in the lab tool and appends ,true if they satisfy the lab constraints and ,false otherwise.
Thx, Jeff! Omei, I was so happy about the two scripts working so well together and being able to see stable versus unstable, that I forgot a step and went straight with the output of Jeffs script. I appologize for the confusion.
You could do this with my BulkLoadV3 booster script. Pick one of the designs as the mutation base, then list the bases that are different in the other design e.g. U15,G20,G32,A24,U25,… Prepend this list with a bang (!) and it will try single and double mutations from the list, or pick a subset of a half-dozen or so and separate them by bangs and it will try all combinations of substitutions, e.g. U15!G20!G32!A24!U25 will try 32 combinations including the original.